Quick start
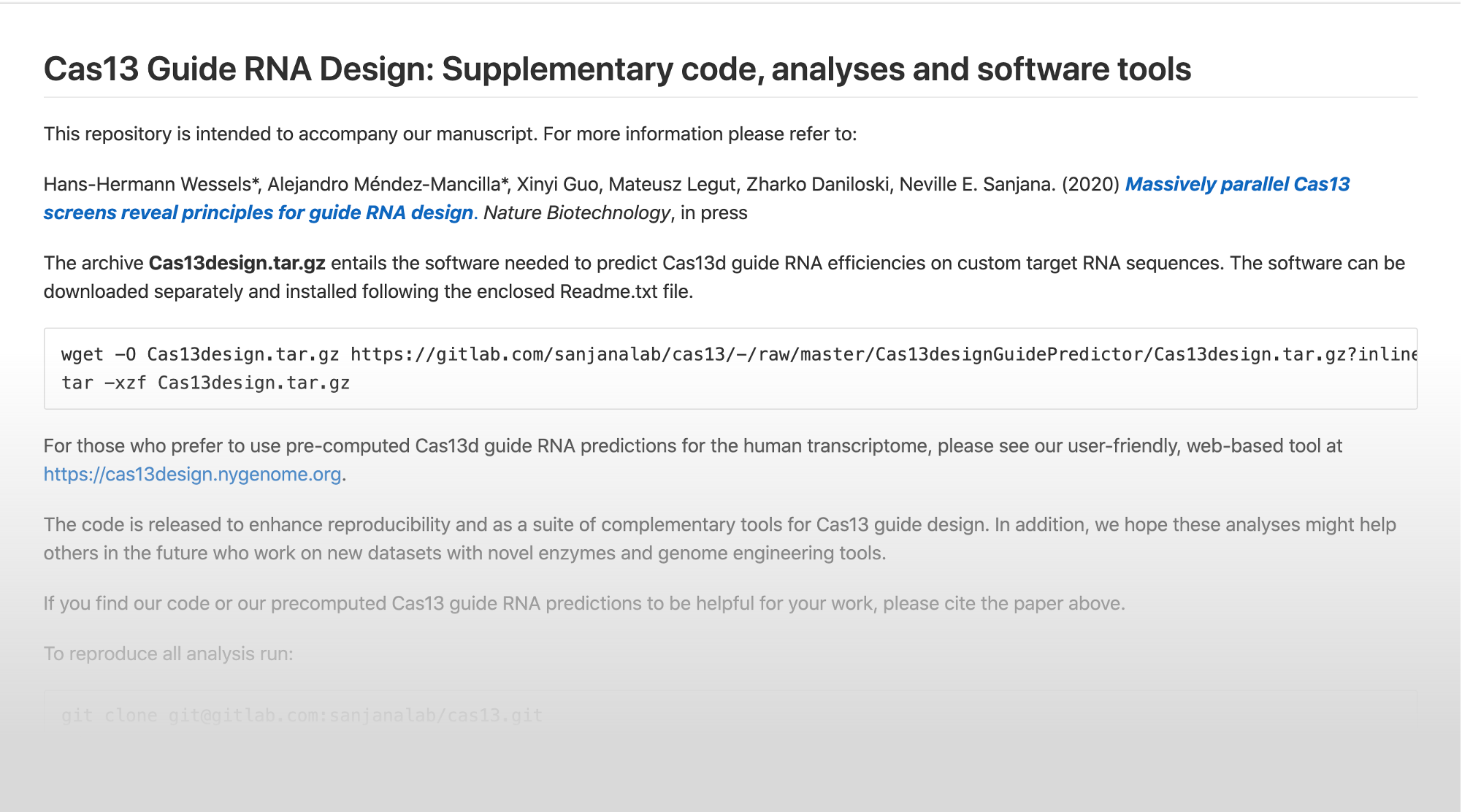
This resource provides optimized guide RNAs to target transcripts in the human transcriptome, model organisms and viral RNA genomes.
This online resource for the prediction Cas13d guide RNAs for target transcripts integrates guide RNA design rules from the following manuscripts:
 Sanjana Lab
Sanjana Lab
Here are the steps to use the design tool:
-
Navigate to the desired species. For human transcript gRNA design, users can (optionally) access a list of all associated Ensembl transcript isoforms for a gene along with expression data for each isoform in different human cell lines by typing the desired gene in the table stored under the Lookup human transcript expression tab.
-
After finding the Ensembl transcript ID of the desired transcript, either click or type it in the search box on the upper left side.
-
Upon lookup, you can browse individual guide RNAs and download a graphical representation of guide RNAs or a table with all guide RNAs that target the transcript.
This online resource for the prediction Cas13d guide RNAs for target transcripts integrates guide RNA design rules from the following manuscripts:
Massively parallel Cas13 screens reveal principles for guide RNA design
Hans-Hermann Wessels*, Alejandro Méndez-Mancilla*, Xinyi Guo, Mateusz Legut, Zharko Daniloski, Neville E. Sanjana
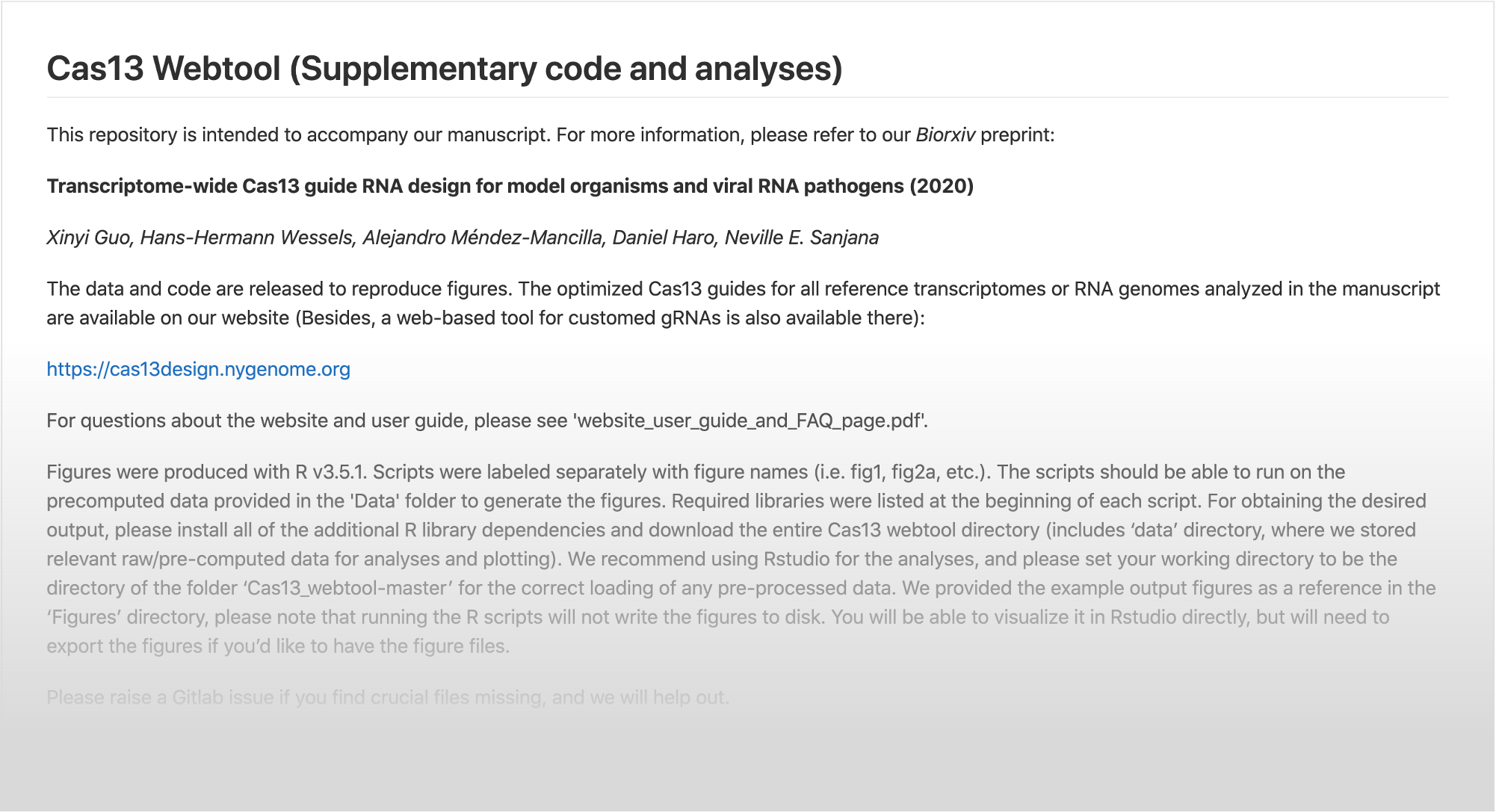
Transcriptome-wide Cas13 guide RNA design for model organisms and viral RNA pathogens
Xinyi Guo, Jahan Rahman, Hans-Hermann Wessels, Alejandro Méndez-Mancilla, Daniel Haro, Xinru Chen, Neville E. Sanjana
* These authors contributed equallyThis resource currently features:
- Selection of perfect match guides for all human protein coding and non-coding genes, separted by transcript and mRNA sub-annotation categories (5'UTR, CDS, 3'UTR).
- Integration of Cancer Cell Line Encyclopedia (CCLE) gene expression data for the selection by cell type specific expression level.
- Selection of perfect match guides for protein coding and non-coding genes in the main model organism, separted by transcript and mRNA sub-annotation categories (5'UTR, CDS, 3'UTR).
- Perfect match guides for SARS-CoV-2, MERS, H1N1 and HIV1 RNA virus genes
Please retrieve transcript isoform expression information directly from the CCLE repository if you don't find your cell line represented.
Release notes:
- 11.2020: v1.1 more user-friendly layout+search, interactive plots, multi-sequence/fasta custom grna design, off-target metrics
- 07.2020: v1.0.1 addition of RNA viruses.
- 06.2020: v1.0 guide RNA efficacy predictions for mRNA and ncRNA in human and model organisms.
- 05.2020: v0.3.1 addition of custom guide predictor.
- 12.2019: v0.3 model update with improved prediction accuracy. The model is now trained on four tiling screen data sets.
- 09.2019: v0.2 improved data structure to allow real-time start up. Adding improved visualizations and data download.
- 07.2019: v0.1 goes online
 Sanjana Lab
Sanjana Lab
Design custom gRNAs
To accurately predict local target site accessibility use sequences of > 80 nt length.
To limit run time when designing guides on webpage, please use target sequences of < 500 nt length. For greater length requirements, please use the 'upload multiple single-entry fasta files' option.
Please be patient while the gRNAs are being scored. (100nt ~10sec; 500nt ~ 30sec)
Please note that the provided fasta file should only contain a single-entry. Otherwise, design results will only be provided for the first sequence in the file.
Please note that all provided fasta files should only contain a single-entry. Otherwise, design results will only be provided for the first sequence in each file.
Cas13 guide RNA predictions
Please use the search field for searching or rank the guide RNA predictions column-wise. Download predictionsLoading...
Lookup human transcript expression

Please use the Search fields to type in a gene symbol or Ensembl gene ID to retrieve its transcript isoforms and their expression in different human cell lines. Transcript expression data is given as transcripts per million (TPM)
Copy the transcript ID and paste it in the submission field in the next Design guide RNAs tab.
Human gRNA design
GENCODE v19 (hg19)
Find your transcript isoform of interest based on CCLE data in the previous tab Lookup transcript.
Cas13 guide RNA predictions along the target transcript
Cas13 guide RNA scores and quartiles are assigned according to the tiling screens used to predict guide efficiency. Download plotCas13 guide RNA predictions
Please use the search field for searching or rank the guide RNA predictions column-wise. Download tableSelect a model organism
M. musculus
GENCODE M24 (mm10)
Cas13 guide RNA predictions along the target transcript
Cas13 guide RNA scores and quartiles are assigned according to the tiling screens used to predict guide efficiency. Download plotCas13 guide RNA predictions
Please use the search field for searching or rank the guide RNA predictions column-wise. Download tableD. rerio
Ensembl v99 (GRCz11)
Cas13 guide RNA predictions along the target transcript
Cas13 guide RNA scores and quartiles are assigned according to the tiling screens used to predict guide efficiency. Download plotCas13 guide RNA predictions
Please use the search field for searching or rank the guide RNA predictions column-wise. Download tableD. melanogaster
Ensembl v99 (BDGP6)
Cas13 guide RNA predictions along the target transcript
Cas13 guide RNA scores and quartiles are assigned according to the tiling screens used to predict guide efficiency. Download plotCas13 guide RNA predictions
Please use the search field for searching or rank the guide RNA predictions column-wise. Download tableC. elegans
Ensembl v99 (WBcel235)
Cas13 guide RNA predictions along the target transcript
Cas13 guide RNA scores and quartiles are assigned according to the tiling screens used to predict guide efficiency. Download plotCas13 guide RNA predictions
Please use the search field for searching or rank the guide RNA predictions column-wise. Download tableA. thaliana
Ensembl Plants v46 (TAIR10)
Cas13 guide RNA predictions along the target transcript
Cas13 guide RNA scores and quartiles are assigned according to the tiling screens used to predict guide efficiency. Download plotCas13 guide RNA predictions
Please use the search field for searching or rank the guide RNA predictions column-wise. Download tableSelect a virus
SARS-CoV-2
Cas13 guide RNA predictions along the target transcript
Cas13 guide RNA scores and quartiles are assigned according to the tiling screens used to predict guide efficiency. Download plotCas13 guide RNA predictions
Please use the search field for searching or rank the guide RNA predictions column-wise. Download tableMERS
Cas13 guide RNA predictions along the target transcript
Cas13 guide RNA scores and quartiles are assigned according to the tiling screens used to predict guide efficiency. Download plotCas13 guide RNA predictions
Please use the search field for searching or rank the guide RNA predictions column-wise. Download tableH1N1
Cas13 guide RNA predictions along the target transcript
Cas13 guide RNA scores and quartiles are assigned according to the tiling screens used to predict guide efficiency. Download plotCas13 guide RNA predictions
Please use the search field for searching or rank the guide RNA predictions column-wise. Download tableHIV1
Cas13 guide RNA predictions along the target transcript
Cas13 guide RNA scores and quartiles are assigned according to the tiling screens used to predict guide efficiency. Download plotCas13 guide RNA predictions
Please use the search field for searching or rank the guide RNA predictions column-wise. Download tableFor large-scale design of Cas13 gRNAs of your own, please refer to our Git archive which contains an installation guide and tutorial for custom reagent design. For supplementary code and analyses of the pre-computed data, please refer to to our other Git archive.
Batch download
Download all designed mRNA/ncRNA guides for single species.
Organisms:
Viruses: